
The vaccines studied were mRNA-based (Moderna mRNA-1273, Pfizer BNT162b2), vector-based (Janssen, AstraZeneca AZD1222, Gamaleya National Center of Epidemiology and Microbiology Sputnik V), protein-based (Novavax NVX-CoV2373), and inactivated virions (Sinopharm BBIP-CorV). 1) for Omicron sublineages after homologous prime/boost vaccination showed low titers and thus significant immune escape when compared to neutralization against ancestral virus Wuhan-Hu-1 (B.1). c The polyclonal plasma neutralizing antibody responses (geometric mean titers, GMT as described in Bowen et al. 
b The S proteins of all Omicron sublineages are less capable to fuse cells compared to the S proteins of Wuhan-Hu-1 or Delta, the latter of which is known to possess high cell-cell fusion capacity. a The RBDs of BA.1, BA.2 and BA.4/5 have an about 2–6-fold higher affinity for ACE2 than the RBDs of the SARS-CoV-2 isolate Wuhan-Hu-1 and the delta variant. Higher binding affinity, lower syncytium formation, and more immune escape by Omicron. But why do the RBDs of Omicron sublineages have high affinities for ACE2 despite harboring several mutations that weaken ACE2 interaction? The answer is provided by another study, which revealed that some Omicron-specific RBD mutations establish new ACE2 contacts and thus compensate for RBD mutations that weaken ACE2 interaction. Interestingly, the RBD of BA.2.12.1, which differs from the BA.2 RBD by only one mutation, has a much lower ACE2 affinity than BA.2 RBD, indicating that this difference is due to mutation L452Q. show that the RBDs of BA.1, BA.2 and BA.4/5 (identical on protein sequence) have a ~2–6-fold higher affinity for ACE2 than the RBDs of the SARS-CoV-2 isolate Wuhan-Hu-1 and the delta variant (Fig. Using biolayer interferometry and surface plasmon resonance analyses, Bowen et al. Thus, one would assume that Omicron sublineages would have reduced ACE2 affinity. While some RBD mutations are known to augment ACE2 binding, most of them weaken spike-ACE2 interaction. 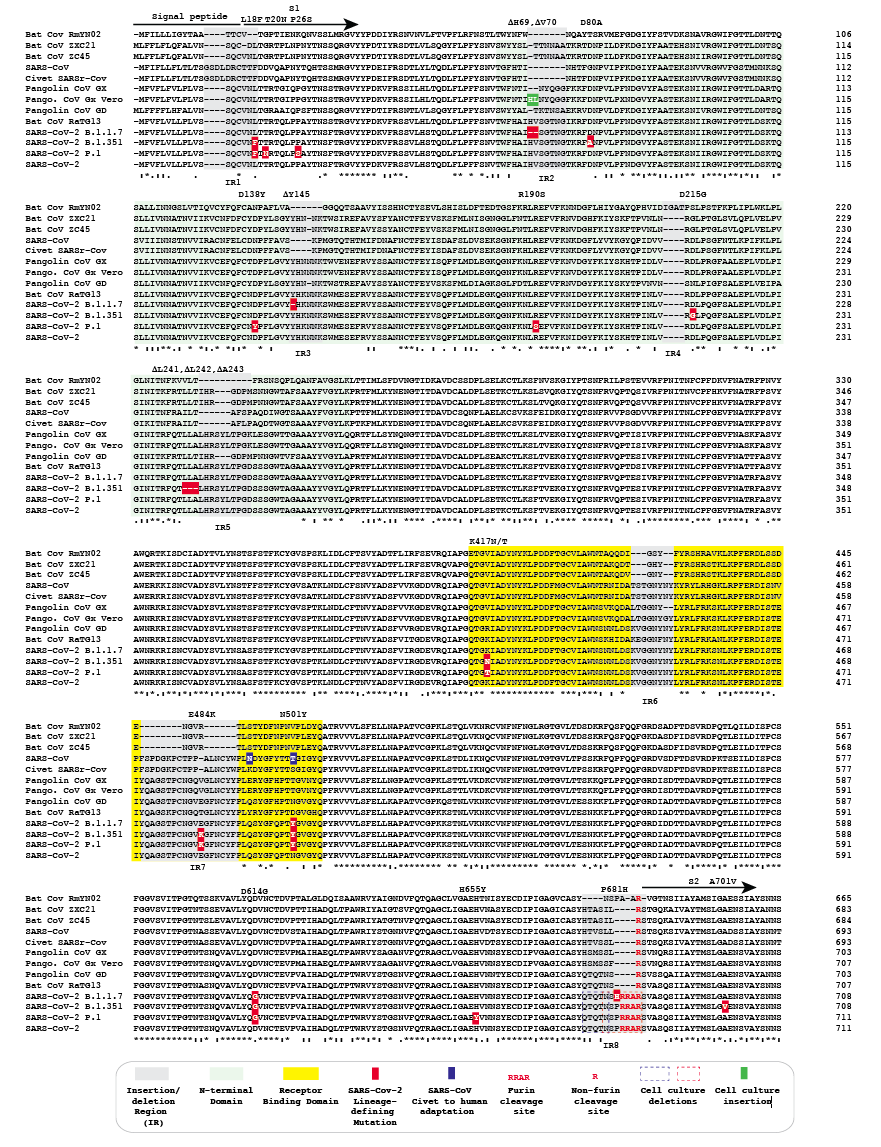
The S proteins of the Omicron sublineages harbor multiple mutations, many of them reside in the RBD and represent escape mutations that reduce the ability of antibody binding. Finally, the SARS-CoV-2 S protein is the main target of neutralizing antibodies, which are correlated for protection against SARS-CoV-2 infection and severe COVID-19. S protein expression on the cell surface leads to fusion with neighboring cells and the formation of giant cells with multiple nuclei, so-called syncytia, which were observed in postmortem biopsies from patients that died from COVID-19. The SARS-CoV-2 S protein also is a pathogenicity factor. Next, the viral genome is released into the cytoplasm, and translation of viral proteins and genome replication start. 2 Subsequently, the S protein is primed by host cell proteases and drives the fusion of the viral and cellular membranes, which can take place at the plasma membrane or within endo-/lysosomes. For SARS-CoV-2 attachment, the S protein binds to the cellular protein angiotensin-converting enzyme 2 (ACE2). The SARS-CoV-2 spike (S) protein is crucial for cell entry, pathogenicity, and is the key target of the adaptive immune response.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
